Visual and Biomedical Datasets the Focus at Second HASTE Data Workshop

What an 1854 map of a London neighborhood can tell us about the current COVID-19 outbreak, and how researchers can use certain datasets to fight the virus were among the thought-provoking ideas presented at the second HASTE Data Workshop.
The workshop, held Wednesday, featured speakers John Lafferty, Smita Krishnaswamy, and Feimei Liu. The series, which is hosted by SEAS Dean and Dean of Science in the Faculty of Arts and Sciences Jeffrey Brock, brings together researchers from across numerous disciplines to present COVID-19-related research and discuss how members of the Yale community can contribute.
 John Lafferty, the John C. Malone Professor of Statistics & Data Science and Computer Science, discussed the importance of interactive visualizations during the COVID-19 crisis, how they can be useful, and some of the pitfalls they can present. Lafferty was one of the developers of the YData course, designed with the idea that all Yale students should learn the skills to use empirical evidence in data to form opinions and make decisions.
John Lafferty, the John C. Malone Professor of Statistics & Data Science and Computer Science, discussed the importance of interactive visualizations during the COVID-19 crisis, how they can be useful, and some of the pitfalls they can present. Lafferty was one of the developers of the YData course, designed with the idea that all Yale students should learn the skills to use empirical evidence in data to form opinions and make decisions.
“And I would say that if ever there was a time when these skills are needed, that time is now,” Lafferty said.
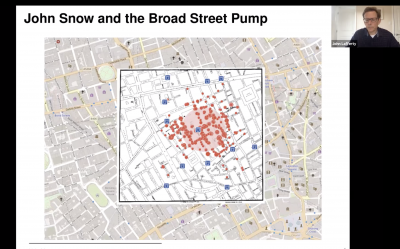
One early example of the impact that data visualizations can have can be found in John Snow’s map of the Broad Street pump in the Soho section of London during the 1854 cholera outbreak. It was commonly believed that the disease was transmitted through the air, but Snow, a physician, posited that it spread through the water. His map helped him make that case to the public.
Lafferty discussed numerous ways that the data being generated about the spread of COVID-19 can be processed to provide similar insights. For instance, a data visualization focusing on the border between Kentucky and Tennessee - two states that have taken very different approaches to shelter-in-place policies - could be the contemporary equivalent to John Snow’s map.
Smita Krishnaswamy, Assistant Professor of Genetics and of Computer Science, gave a detailed look at how the coronavirus infects the body, how the body fights it - and the ways that the body’s own immune system can add to the trouble. 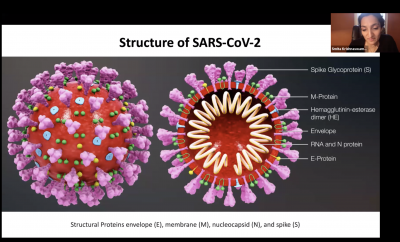
To design a vaccine, she noted, scientists are asking questions such as how mutations affect the ability of the virus to infect, and what kinds of antibodies are good at neutralizing the virus. Certain types of datasets, such as those related to viral genome sequencing, single cell RNA sequencing, and protein-nanobody binding can help answer those questions, she said.
Regarding protein-nanobody binding, she noted that the laboratory of Yale immunobiologist Aaron Ring has generated a massive dataset of proteins with a large library of nanobodies, showing which proteins bind to which nanobodies.
“Which is fantastic for training a type of machine learning or deep learning for this kind of thing,” she said.
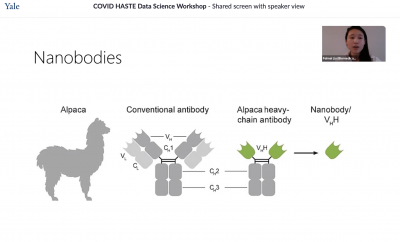 Feimei Liu, a Ph.D. candidate in biomedical engineering, also discussed protein-nanobody binding. She explained the multi-step process of nanobody discovery, which often involves immunizing alpacas and can take at least four months.
Feimei Liu, a Ph.D. candidate in biomedical engineering, also discussed protein-nanobody binding. She explained the multi-step process of nanobody discovery, which often involves immunizing alpacas and can take at least four months.
“It's a lot of work and agony for discovering one good nanobody,” she said.
To make things easier, she said, Yale has teamed up with researchers at Harvard and the University of California, San Francisco to design a synthetic nanobody library that mimics the natural repertoire of nanobodies. They now have 24 billion different nanobodies. The next goal, she said, is to discover nanobodies for all human antigens so they can share the sequences of all nanobodies for everyone in the community to use for their own research.
The next HASTE workshop takes place Tuesday, April 21, 4-5 p.m.

