A New Focus on Proteins
The average family photo, all toothy grins and warm embraces, doesn't pass scientific muster. How many strands of hair are on Dad's head? Was it an ill-timed sneeze that turned the toddler into a blurry kangaroo?
Structural biologists face this same predicament. Though the high-impact proteins they most want to study can be seen with an electron microscope, the resulting pictures are "noisy." Even viewed at movement-reducing cryogenic temperatures (a technique known as electron cryo-microscopy, or cryo-EM), the proteins' structures are often obscured by scurrying electrons, alignment errors, and extraneous particles.
 To separate the structure from the noise, scientists must measure how much each part of the structure is "resolved," a task typically—and problematically—done using the Fourier Shell Correlation (FSC) procedure, which results in a one-size-fits-all resolution. This global resolution is similar to a camera lens that can bring into focus only the foreground or the background.
To separate the structure from the noise, scientists must measure how much each part of the structure is "resolved," a task typically—and problematically—done using the Fourier Shell Correlation (FSC) procedure, which results in a one-size-fits-all resolution. This global resolution is similar to a camera lens that can bring into focus only the foreground or the background.
To address this limitation, a team of Yale engineers, consisting of Professor of Diagnostic Radiology, Electrical Engineering, & Biomedical Engineering Hemant Tagare, Professor of Physiology & Biomedical Engineering Fred Sigworth, and doctoral student Alp Kucukelbir, have proposed a mathematical theory for measuring local resolution for each part of the structure. The team's novel and statistically-rigorous approach adjusts the resolution based on local sinusoidal features observed for each voxel, then correlates and filters the data using likelihood-ratio tests and a false discovery rate procedure to ensure accuracy.
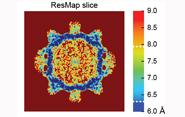
The team's theory, recently published in Nature Methods, has also been developed by Kucukelbir into an efficient publicly-accessible algorithm known as ResMap. Using previously published density maps, the team's article shows that ResMap results are comparable to or better than other FSC plots—with the added flexibility of local resolution. Additionally, ResMap images can be quickly produced even on a desktop computer.
In future research, the team looks to determine why the resolution changes in the first place, a "focus" that ensures a better picture of proteins.

